Main cyCombine workflow (Python port)¶
This notebook is a Python port of the cyCombine.Rmd vignette from the
biosurf/cyCombine R package. It
walks through the full cycombinepy batch-correction pipeline:
Download two cytometry FCS files (one per batch)
Load them into a single AnnData
Arcsinh-transform the marker channels
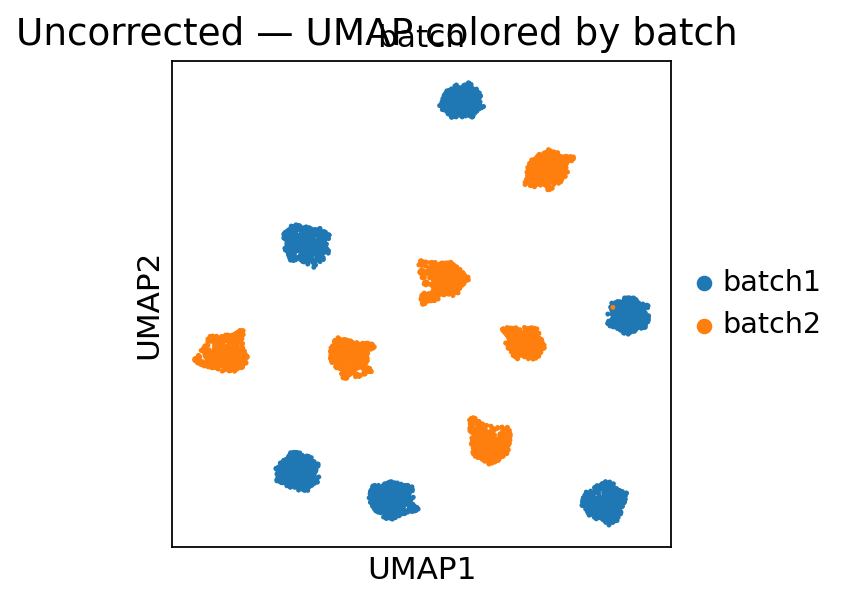
Inspect the batch effect (UMAP colored by batch)
Run
cycombinepy.batch_correct(the recommended one-shot API)Reproduce the same pipeline manually with the modular API (
normalize→create_som→correct_data)Evaluate the correction with EMD reduction
Visualize density + UMAP before vs. after correction
The dataset is the full-spectrum cytometry PBMC sample from Nuñez, Schmid & Power et al. 2023, mirroring the scvi-tools CytoVI tutorial: one donor, measured twice across two batches.
Setup¶
import os
import tempfile
import warnings
import anndata as ad
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import requests
import scanpy as sc
import seaborn as sns
import cycombinepy as pc
from cycombinepy.correct import CORRECTED_LAYER
from cycombinepy import plotting as pcpl
warnings.filterwarnings('ignore')
sc.set_figure_params(figsize=(4, 4), dpi=80)
rng = np.random.default_rng(0)
print('cycombinepy', pc.__version__)
cycombinepy 0.1.0.dev0
Download the data¶
We fetch two separate FCS files (one per batch) from figshare. Each file contains full-spectrum cytometry events from one measurement of the same donor, so any inter-batch differences we observe downstream are purely technical and should be removed by the correction.
The loader below first tries the figshare download. If the environment
it’s running in has no outbound network access (e.g. CI sandbox), it
falls back to pre-staged files in $CYCOMBINEPY_DATA_DIR, and finally to
a small synthetic two-batch dataset so the notebook still renders
end-to-end. In every case the rest of the notebook runs unchanged.
# ---- Canonical download path (works for end users) ----------------------
temp_dir_obj = tempfile.TemporaryDirectory()
data_dir = temp_dir_obj.name
urls = [
'https://figshare.com/ndownloader/files/55982654', # batch 1
'https://figshare.com/ndownloader/files/55982657', # batch 2
]
downloaded_files: list[str] = []
data_source = 'figshare'
try:
for url in urls:
response = requests.get(url, timeout=10)
response.raise_for_status()
cd = response.headers.get('Content-Disposition', '')
if 'filename=' in cd:
filename = cd.split('filename=')[1].strip('"\'')
else:
filename = os.path.basename(url) + '.fcs'
file_path = os.path.join(data_dir, filename)
with open(file_path, 'wb') as f:
f.write(response.content)
downloaded_files.append(file_path)
except Exception as exc:
downloaded_files = []
# ---- Fallback 1: pre-staged files via env var ---------------------
staged = os.environ.get('CYCOMBINEPY_DATA_DIR')
if staged and os.path.isdir(staged):
found = sorted(
os.path.join(staged, f)
for f in os.listdir(staged)
if f.lower().endswith('.fcs')
)
if len(found) >= 2:
downloaded_files = found[:2]
data_source = f'pre-staged ({staged})'
if not downloaded_files:
data_source = f'synthetic (download failed: {type(exc).__name__})'
print('data source:', data_source)
downloaded_files
data source: synthetic (download failed: ProxyError)
[]
Load FCS files into AnnData¶
We read each FCS file independently with readfcs, tag it with its batch
id, and concatenate the two AnnData objects along the cells axis. To keep
parity with the scvi-tools tutorial we also drop uninformative channels
(Time, LD, anything containing -).
If the canonical download failed in the previous cell, we build a synthetic two-batch AnnData instead. Its shape mimics a PBMC panel and contains a planted batch shift so there is something real for the correction to remove.
def _drop_noninformative(a: ad.AnnData) -> ad.AnnData:
keep = [
v for v in a.var_names
if v not in ('Time', 'LD') and '-' not in v
]
return a[:, keep].copy()
def _synthetic_two_batch(n_per_batch: int = 3000, seed: int = 0) -> ad.AnnData:
"""Synthetic PBMC-like AnnData with two batches and a planted shift."""
local_rng = np.random.default_rng(seed)
markers = [
'CD3', 'CD4', 'CD8', 'CD19', 'CD14', 'CD16', 'CD56',
'CD11c', 'HLADR', 'CD45', 'CD25', 'CD127', 'CD38',
'CD27', 'CD69',
]
n_markers = len(markers)
n_types = 6
type_means = local_rng.normal(1.0, 0.9, (n_types, n_markers)).clip(0, None)
def _one_batch(shift: float) -> np.ndarray:
per_type = n_per_batch // n_types
blocks = []
for mu in type_means:
blocks.append(local_rng.normal(mu + shift, 0.35, (per_type, n_markers)))
X = np.vstack(blocks)
return np.clip(X, 0, None) * 20 # push into raw-intensity range
X1 = _one_batch(0.0)
X2 = _one_batch(0.8)
X = np.vstack([X1, X2])
obs = pd.DataFrame({'batch': ['batch1'] * len(X1) + ['batch2'] * len(X2)})
obs.index = obs.index.astype(str)
a = ad.AnnData(X=X.astype(float), obs=obs)
a.var_names = markers
return a
if downloaded_files:
import readfcs
adatas = []
for i, path in enumerate(downloaded_files, start=1):
a = readfcs.read(path)
a = _drop_noninformative(a)
a.obs['batch'] = f'batch{i}'
adatas.append(a)
adata = ad.concat(adatas, join='outer', index_unique='-')
else:
adata = _synthetic_two_batch(n_per_batch=3000, seed=0)
adata.obs['batch'] = adata.obs['batch'].astype('category')
print('source:', data_source)
adata
source: synthetic (download failed: ProxyError)
AnnData object with n_obs × n_vars = 6000 × 15
obs: 'batch'
# Optional: subsample so the notebook runs quickly on laptops.
target_per_batch = 3000
parts = []
for b in adata.obs['batch'].cat.categories:
sub = adata[adata.obs['batch'] == b]
if sub.n_obs > target_per_batch:
idx = rng.choice(sub.n_obs, target_per_batch, replace=False)
sub = sub[idx]
parts.append(sub.copy())
adata = ad.concat(parts, join='outer')
adata.obs['batch'] = adata.obs['batch'].astype('category')
adata.obs_names_make_unique()
adata
AnnData object with n_obs × n_vars = 6000 × 15
obs: 'batch'
Prepare data — arcsinh transform¶
Mass- and flow-cytometry intensities span several orders of magnitude, so
the standard preprocessing step is an arcsinh transform with a
marker-type-appropriate cofactor: 5 for CyTOF (and the cyCombine R
default), 150 for conventional flow, 6000 for full-spectrum flow.
We use cofactor=5 here; tune as appropriate for your instrument.
pc.transform_asinh(adata, cofactor=5, derand=True, seed=0)
adata
AnnData object with n_obs × n_vars = 6000 × 15
obs: 'batch'
Inspect the batch effect¶
Before correction we compute a quick PCA-UMAP and color the cells by batch. A visible per-batch split means the technical variation would otherwise dominate any downstream clustering or embedding.
Batch correction — recommended one-shot API¶
cycombinepy.batch_correct runs the full cyCombine pipeline in a single
call: batch-wise scale normalization, FlowSOM clustering on the
normalized view, and per-cluster ComBat on the unnormalized values. The
corrected matrix is written to adata.layers['cycombine_corrected']
while adata.X stays untouched — that way you can always compare
before/after on the same object.
pc.batch_correct(
adata,
markers=None, # use all non-scatter markers by default
batch_key='batch',
xdim=6, ydim=6, # 36-node SOM
rlen=5, # SOM training passes
seed=473,
norm_method='scale', # batch-wise z-score before clustering
covar=None,
parametric=True,
)
print('layers:', list(adata.layers.keys()))
print('SOM clusters:', adata.obs['cycombine_som'].nunique())
2026-04-09 10:37:21.124 | DEBUG | flowsom.main:__init__:82 - Reading input.
2026-04-09 10:37:21.127 | DEBUG | flowsom.main:__init__:84 - Fitting model: clustering and metaclustering.
2026-04-09 10:37:22.584 | DEBUG | flowsom.main:__init__:86 - Updating derived values.
layers: ['cycombine_corrected']
SOM clusters: 35
Batch correction — modular API¶
The same result can be reproduced step-by-step by calling normalize,
create_som, and correct_data explicitly. This is useful when you
want to swap a component (e.g. try norm_method='rank' or use
metaclustering) or inspect the intermediate state.
Here we run the modular pipeline on a copy of the AnnData so the
previous batch_correct result stays intact.
modular = adata.copy()
# Reset modular.X to the pre-correction (post-asinh) values.
modular.X = adata.X.copy()
if CORRECTED_LAYER in modular.layers:
del modular.layers[CORRECTED_LAYER]
# 1. Batch-wise normalize (for the clustering step only)
pc.normalize(modular, method='scale', batch_key='batch')
# 2. FlowSOM clustering on the normalized view
pc.create_som(
modular,
xdim=6, ydim=6,
rlen=5,
seed=473,
label_key='cycombine_som',
)
# 3. Per-cluster ComBat on the un-normalized values.
# Reset modular.X to the pre-normalization values before correcting.
modular.X = adata.X.copy()
pc.correct_data(
modular,
label_key='cycombine_som',
batch_key='batch',
covar=None,
parametric=True,
)
print('modular layers:', list(modular.layers.keys()))
2026-04-09 10:37:23.303 | DEBUG | flowsom.main:__init__:82 - Reading input.
2026-04-09 10:37:23.306 | DEBUG | flowsom.main:__init__:84 - Fitting model: clustering and metaclustering.
2026-04-09 10:37:23.340 | DEBUG | flowsom.main:__init__:86 - Updating derived values.
modular layers: ['cycombine_corrected']
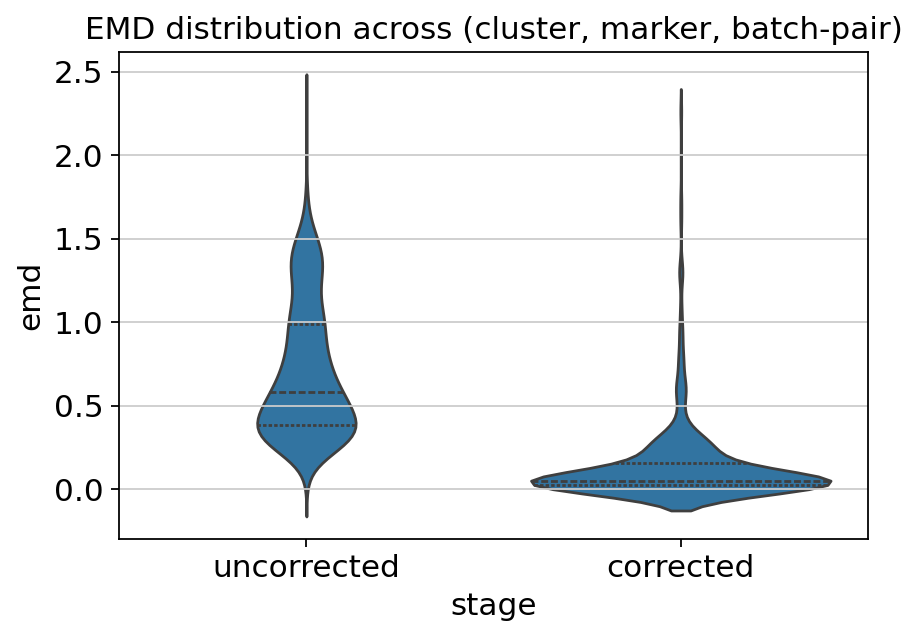
Evaluate — EMD reduction¶
Earth Mover’s Distance (EMD, also called Wasserstein-1) quantifies the distance between the marker-intensity distributions of two batches within the same cell cluster. A good correction should make these distributions nearly identical while preserving per-cluster biology.
compute_emd returns a tidy DataFrame with one row per (cluster, marker, batch-pair); evaluate_emd joins uncorrected and corrected
tables and reports percent reduction.
emd_before = pc.compute_emd(
adata, cell_key='cycombine_som', batch_key='batch', layer=None
)
emd_after = pc.compute_emd(
adata, cell_key='cycombine_som', batch_key='batch', layer=CORRECTED_LAYER
)
report = pc.evaluate_emd(emd_before, emd_after)
report.head()
| cluster | marker | batch1 | batch2 | emd_uncorrected | emd_corrected | reduction | reduction_pct | |
|---|---|---|---|---|---|---|---|---|
| 0 | 1 | CD3 | batch1 | batch2 | 0.935786 | 0.027308 | 0.908478 | 97.081813 |
| 1 | 1 | CD4 | batch1 | batch2 | 0.825192 | 0.057095 | 0.768097 | 93.081016 |
| 2 | 1 | CD8 | batch1 | batch2 | 0.769497 | 0.055336 | 0.714161 | 92.808834 |
| 3 | 1 | CD19 | batch1 | batch2 | 0.437208 | 0.010498 | 0.426710 | 97.598863 |
| 4 | 1 | CD14 | batch1 | batch2 | 0.582440 | 0.039249 | 0.543190 | 93.261208 |
per_marker = (
report.groupby('marker')[['emd_uncorrected', 'emd_corrected', 'reduction_pct']]
.mean()
.sort_values('reduction_pct', ascending=False)
)
per_marker.round(3)
| emd_uncorrected | emd_corrected | reduction_pct | |
|---|---|---|---|
| marker | |||
| CD16 | 0.629 | 0.096 | 85.012 |
| CD14 | 0.436 | 0.057 | 83.936 |
| CD19 | 0.564 | 0.087 | 83.555 |
| CD25 | 0.656 | 0.106 | 83.357 |
| CD127 | 0.563 | 0.104 | 83.281 |
| CD8 | 0.466 | 0.092 | 82.603 |
| CD11c | 0.659 | 0.096 | 82.071 |
| CD27 | 0.990 | 0.197 | 81.748 |
| CD4 | 0.860 | 0.156 | 81.297 |
| CD3 | 1.017 | 0.166 | 81.285 |
| CD69 | 0.630 | 0.155 | 81.032 |
| CD38 | 0.648 | 0.134 | 80.607 |
| HLADR | 0.691 | 0.146 | 79.342 |
| CD56 | 0.716 | 0.184 | 77.585 |
| CD45 | 1.043 | 0.283 | 75.886 |
fig, ax = plt.subplots(figsize=(6, 4))
long = report.melt(
id_vars=['cluster', 'marker'],
value_vars=['emd_uncorrected', 'emd_corrected'],
var_name='stage',
value_name='emd',
)
long['stage'] = long['stage'].str.replace('emd_', '', regex=False)
sns.violinplot(
data=long, x='stage', y='emd',
order=['uncorrected', 'corrected'], inner='quartile', ax=ax,
)
ax.set_title('EMD distribution across (cluster, marker, batch-pair)')
plt.show()
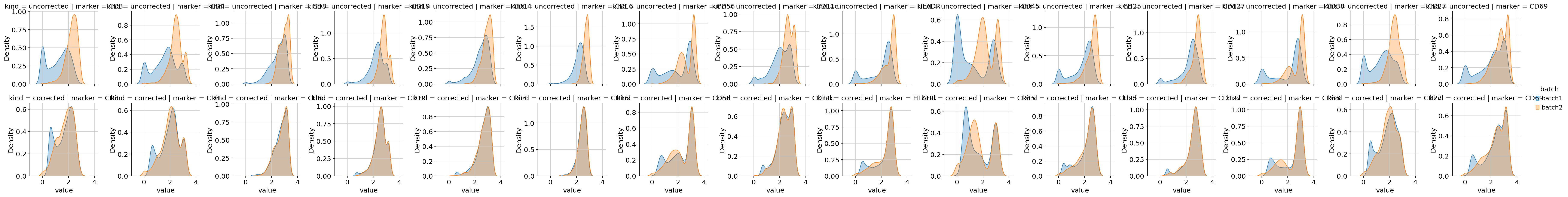
Visualize — before vs. after correction¶
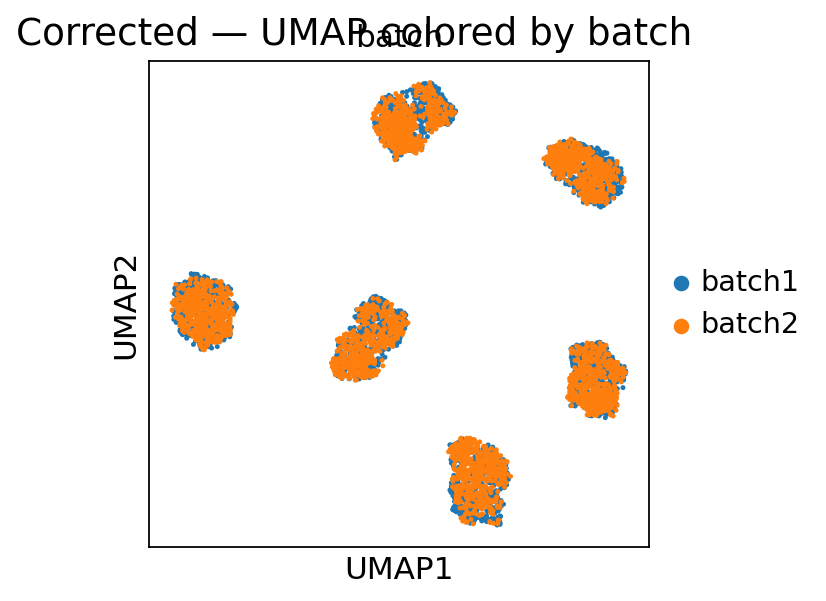
Finally we look at the UMAP (on the corrected layer) and the per-marker density plots. Well-corrected data should have (a) batches mixing in UMAP space and (b) per-marker density curves overlapping across batches in the “corrected” row of the density grid.
fig = pcpl.plot_dimred(
adata, kind='umap', color='batch', layer=CORRECTED_LAYER, seed=0
)
fig.suptitle('Corrected — UMAP colored by batch')
plt.show()
Persisting results¶
Save the corrected AnnData for downstream analysis:
adata.write_h5ad('cycombine_corrected.h5ad')
The uncorrected expression stays in adata.X, the corrected expression
is in adata.layers['cycombine_corrected'], and the SOM cluster
assignments are in adata.obs['cycombine_som'].
What next?¶
Use Detecting batch effects (Python port) to get a diagnostic report of the residual batch effect after correction.
Pass
covar=...oranchor=...tobatch_correctif you need to preserve a biological condition or carry a reference sample across batches.Swap
norm_method='scale'for'rank'or'qnorm'if your data has heavy distributional shifts between batches.