Detecting batch effects (Python port)¶
This notebook is a Python port of the detect_batch_effects.Rmd
vignette from biosurf/cyCombine.
It demonstrates the two batch-effect diagnostics available in
cycombinepy:
detect_batch_effect_express— a fast 3-plot summary: a mean-EMD heatmap across markers, per-marker density curves colored by batch, and an MDS plot of per-sample medians.detect_batch_effect— the above, plus a UMAP of the uncorrected data and a per-marker MAD bar chart.
Run these before you decide to correct: if your data already mixes cleanly across batches, cyCombine correction may do more harm than good.
Cofactor guidance¶
The arcsinh transform compresses the dynamic range of cytometry intensities. Use the cofactor that matches your instrument:
Technology |
Cofactor |
|---|---|
Mass cytometry |
5 |
Conventional flow |
150 |
Full-spectrum flow |
6000 |
Setup¶
import os
import tempfile
import warnings
import anndata as ad
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import requests
import scanpy as sc
import cycombinepy as pc
warnings.filterwarnings('ignore')
sc.set_figure_params(figsize=(4, 4), dpi=80)
print('cycombinepy', pc.__version__)
cycombinepy 0.1.0.dev0
Load the Nuñez PBMC dataset¶
We reuse the two figshare-hosted FCS files from the
Main cyCombine workflow (Python port) tutorial. The loader below tries the canonical
download first, falls back to pre-staged files in
$CYCOMBINEPY_DATA_DIR, and finally synthesizes a two-batch AnnData so
the notebook executes even in offline environments.
def _drop_noninformative(a: ad.AnnData) -> ad.AnnData:
keep = [
v for v in a.var_names
if v not in ('Time', 'LD') and '-' not in v
]
return a[:, keep].copy()
def _synthetic_two_batch(n_per_batch: int = 3000, seed: int = 0) -> ad.AnnData:
local_rng = np.random.default_rng(seed)
markers = [
'CD3', 'CD4', 'CD8', 'CD19', 'CD14', 'CD16', 'CD56',
'CD11c', 'HLADR', 'CD45', 'CD25', 'CD127', 'CD38',
'CD27', 'CD69',
]
n_markers = len(markers)
n_types = 6
type_means = local_rng.normal(1.0, 0.9, (n_types, n_markers)).clip(0, None)
def _one_batch(shift: float) -> np.ndarray:
per_type = n_per_batch // n_types
blocks = []
for mu in type_means:
blocks.append(local_rng.normal(mu + shift, 0.35, (per_type, n_markers)))
return np.clip(np.vstack(blocks), 0, None) * 20
X1 = _one_batch(0.0)
X2 = _one_batch(0.8)
X = np.vstack([X1, X2])
obs = pd.DataFrame({
'batch': ['batch1'] * len(X1) + ['batch2'] * len(X2),
'sample': ['b1_s1'] * len(X1) + ['b2_s1'] * len(X2),
})
obs.index = obs.index.astype(str)
a = ad.AnnData(X=X.astype(float), obs=obs)
a.var_names = markers
return a
def load_pbmc_two_batches() -> tuple[ad.AnnData, str]:
# ---- Canonical figshare download ---------------------------------
temp_dir_obj = tempfile.TemporaryDirectory()
data_dir = temp_dir_obj.name
urls = [
'https://figshare.com/ndownloader/files/55982654',
'https://figshare.com/ndownloader/files/55982657',
]
downloaded: list[str] = []
try:
for url in urls:
r = requests.get(url, timeout=10)
r.raise_for_status()
cd = r.headers.get('Content-Disposition', '')
name = (
cd.split('filename=')[1].strip('"\'')
if 'filename=' in cd
else os.path.basename(url) + '.fcs'
)
path = os.path.join(data_dir, name)
with open(path, 'wb') as f:
f.write(r.content)
downloaded.append(path)
source = 'figshare'
except Exception as exc:
downloaded = []
source = f'synthetic (download failed: {type(exc).__name__})'
staged = os.environ.get('CYCOMBINEPY_DATA_DIR')
if staged and os.path.isdir(staged):
found = sorted(
os.path.join(staged, f)
for f in os.listdir(staged)
if f.lower().endswith('.fcs')
)
if len(found) >= 2:
downloaded = found[:2]
source = f'pre-staged ({staged})'
if downloaded:
import readfcs
adatas = []
for i, path in enumerate(downloaded, start=1):
a = readfcs.read(path)
a = _drop_noninformative(a)
a.obs['batch'] = f'batch{i}'
a.obs['sample'] = f'b{i}_s1'
adatas.append(a)
adata = ad.concat(adatas, join='outer', index_unique='-')
# Keep a handle on the temp dir for this session.
adata.uns['_temp_dir_obj_ref'] = id(temp_dir_obj)
else:
adata = _synthetic_two_batch(n_per_batch=3000, seed=0)
adata.obs['batch'] = adata.obs['batch'].astype('category')
adata.obs['sample'] = adata.obs['sample'].astype('category')
adata.obs_names_make_unique()
return adata, source
adata, data_source = load_pbmc_two_batches()
print('data source:', data_source)
adata
data source: synthetic (download failed: ProxyError)
AnnData object with n_obs × n_vars = 6000 × 15
obs: 'batch', 'sample'
# Arcsinh-transform before diagnostics. cyCombine recommends cofactor 5
# for CyTOF; change to 150 (flow) or 6000 (spectral) as appropriate.
pc.transform_asinh(adata, cofactor=5, derand=True, seed=0)
adata
AnnData object with n_obs × n_vars = 6000 × 15
obs: 'batch', 'sample'
detect_batch_effect_express — the quick summary¶
Call detect_batch_effect_express to get a fast three-panel
visualization of batch imbalance:
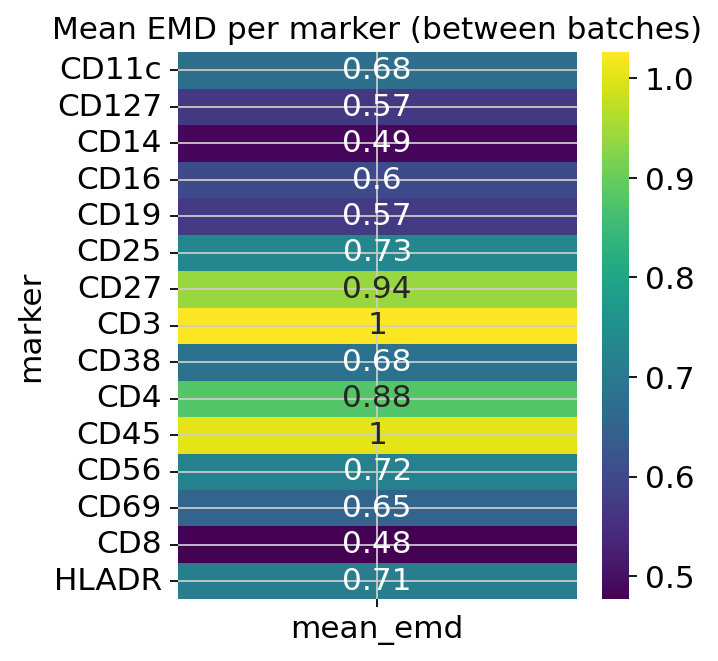
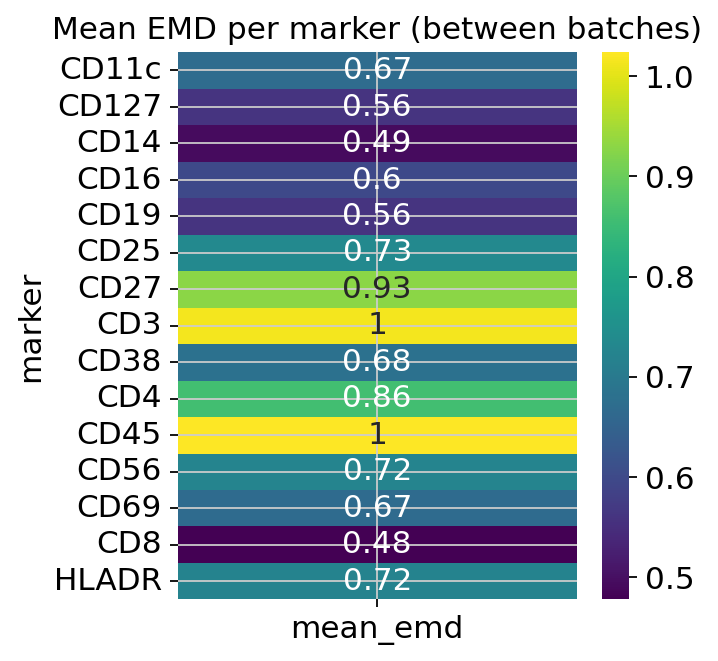
an EMD heatmap of mean Earth Mover’s Distance per marker across all batch pairs,
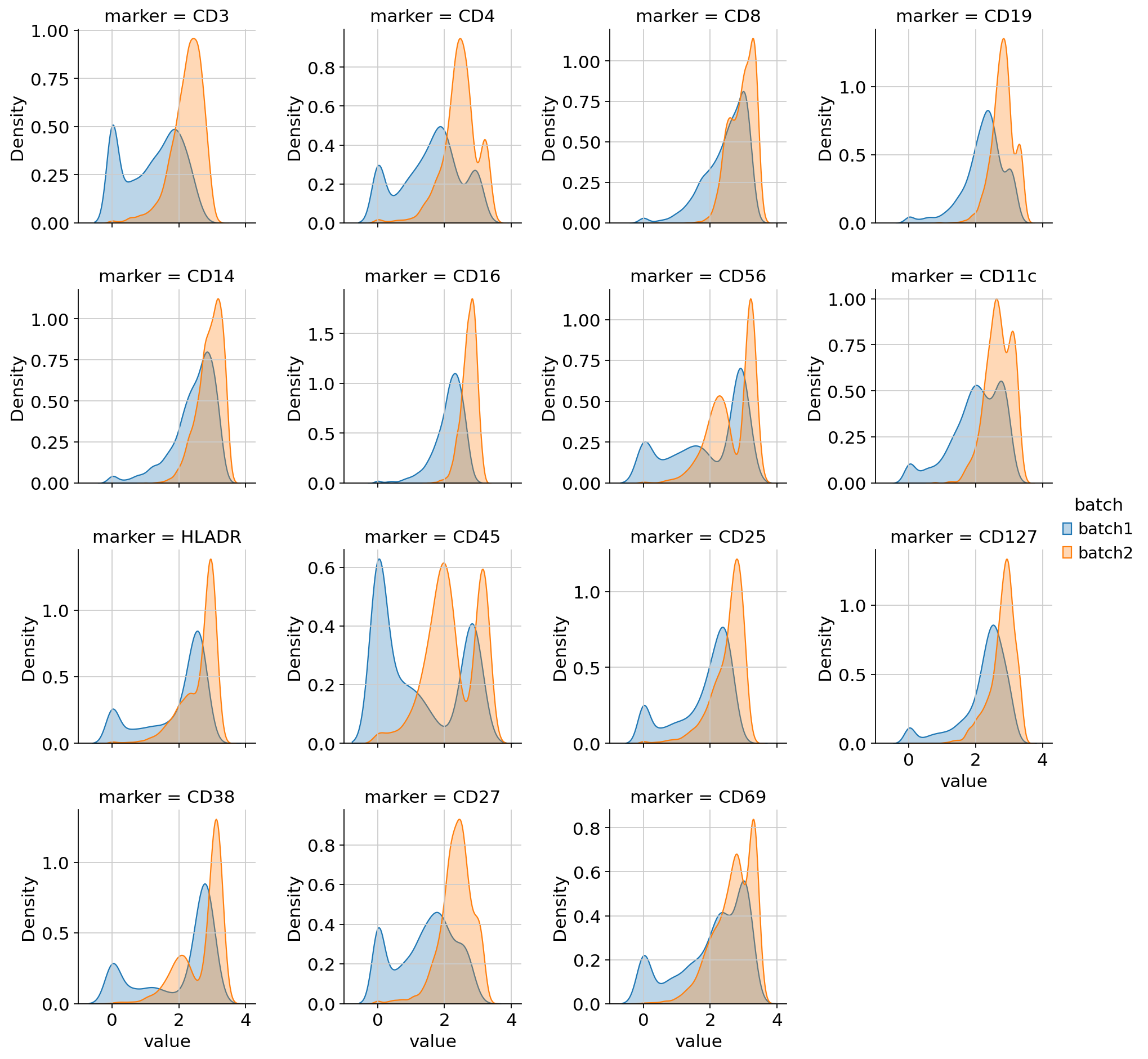
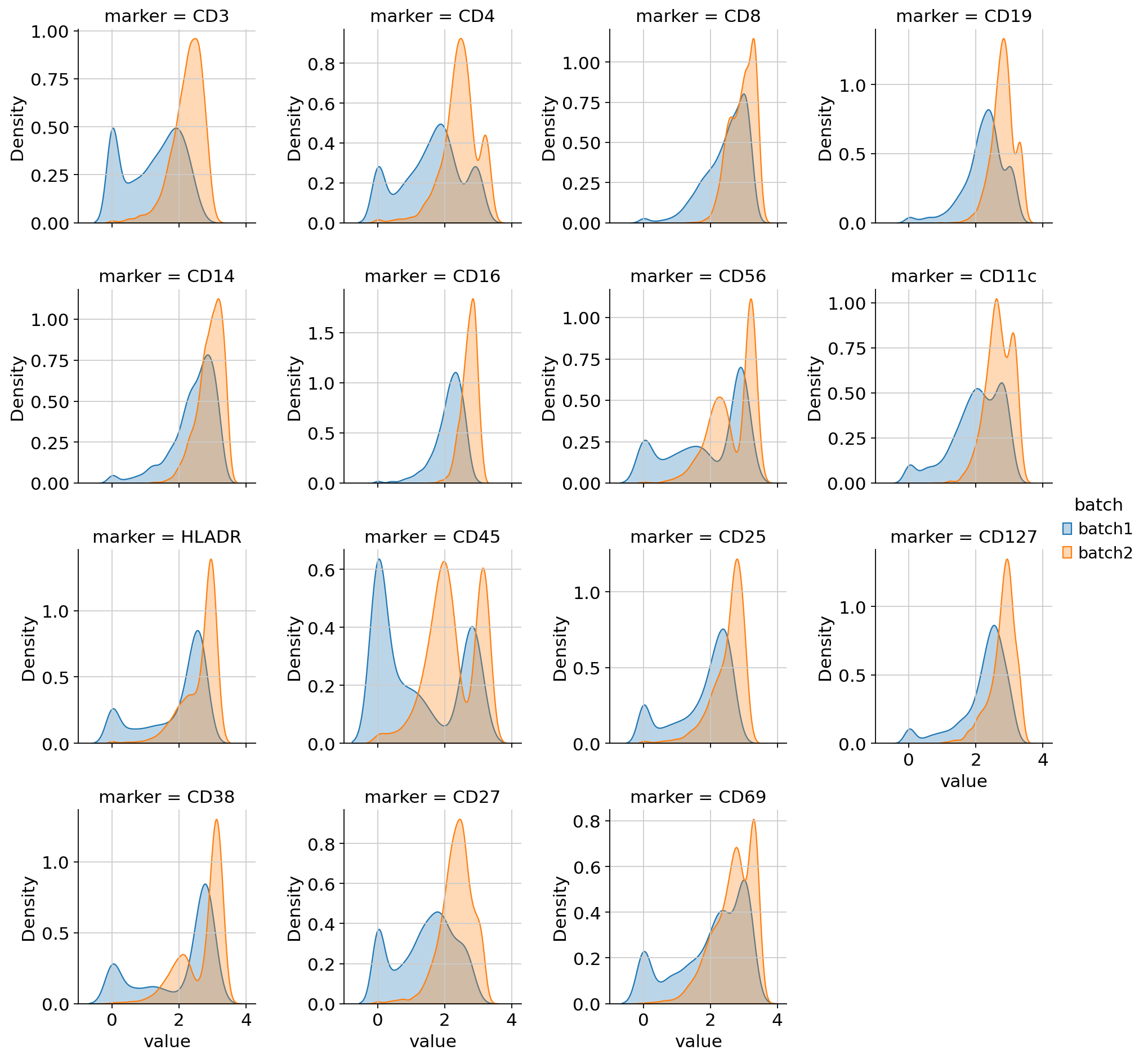
per-marker density curves colored by batch,
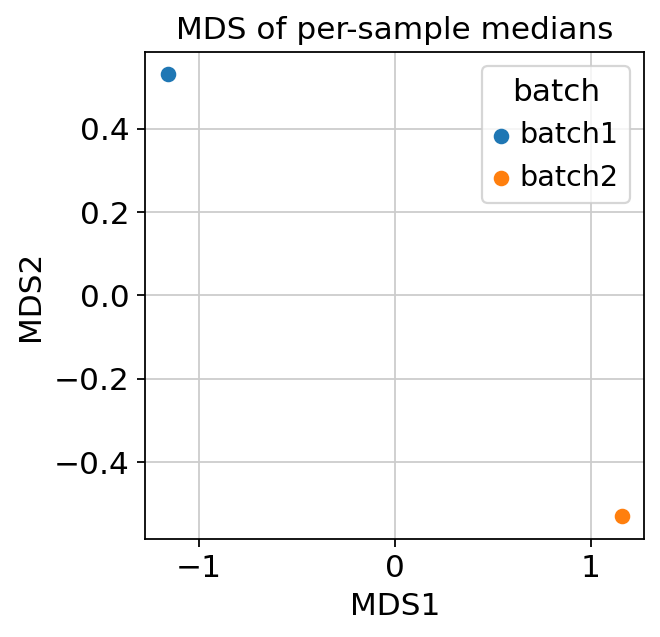
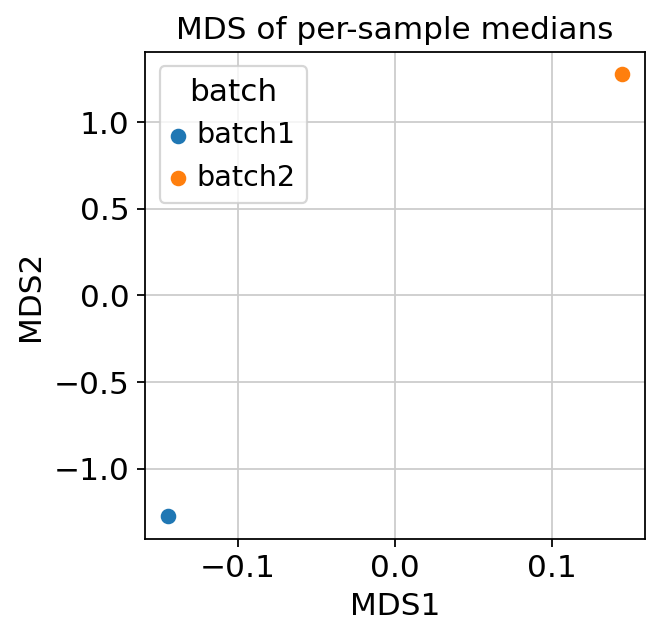
and an MDS plot of per-sample medians (requires
sample_key).
Pass out_dir=... to save all three as PNGs; otherwise the function
returns a dict of matplotlib figures you can display inline.
figs_express = pc.detect_batch_effect_express(
adata,
batch_key='batch',
sample_key='sample',
downsample=5000,
out_dir=None,
seed=472,
)
figs_express['emd']
detect_batch_effect — the full diagnostic¶
detect_batch_effect runs the express diagnostics above and adds a
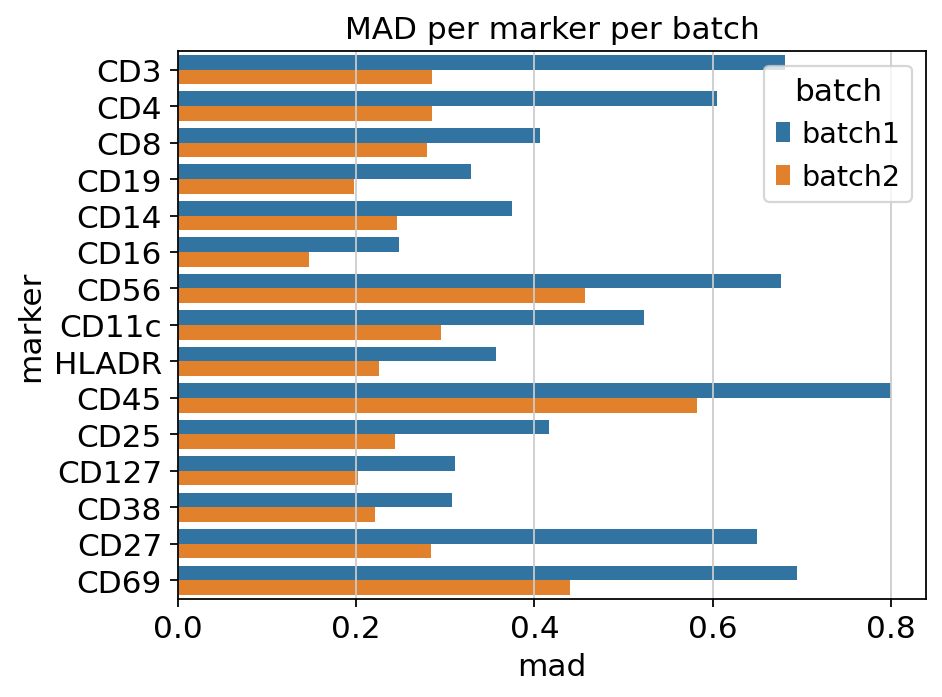
PCA-UMAP colored by batch plus a per-marker MAD bar chart. Use this
when you want a single report to decide whether correction is
warranted.
figs_full = pc.detect_batch_effect(
adata,
batch_key='batch',
sample_key='sample',
downsample=5000,
out_dir=None,
seed=434,
)
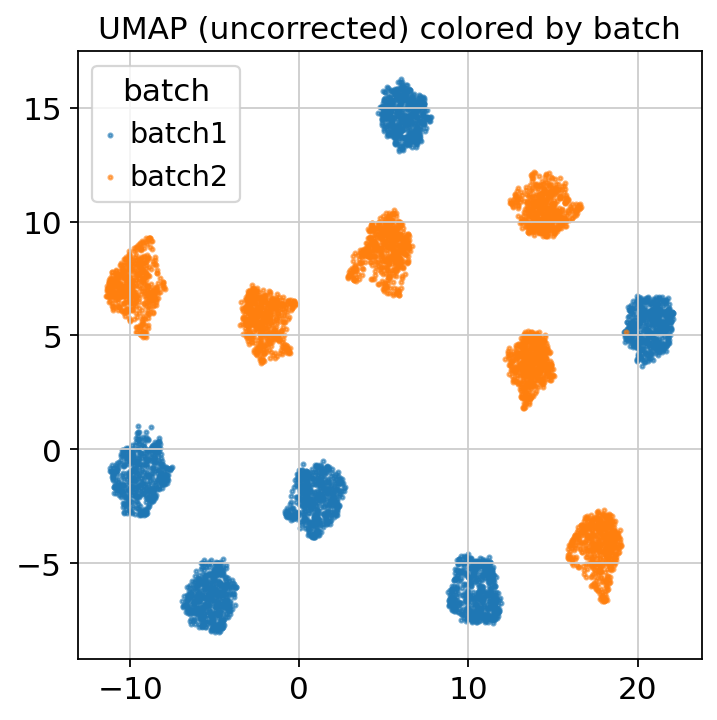
figs_full['umap']
Interpreting the diagnostics¶
EMD heatmap — large per-marker values indicate markers whose distributions differ most across batches. A marker with near-zero EMD is already well-aligned and does not need correction.
Density plots — overlapping curves across batches are good; a bimodal split by batch indicates technical variation.
MDS of per-sample medians — samples clustering by batch rather than by biology is a red flag.
UMAP colored by batch — a clean batch-wise separation means a downstream embedding/clustering will be dominated by technical variation unless you correct.
Per-marker MAD — large per-batch dispersion differences hint at instrument drift or sample-prep variability.
If these diagnostics flag a batch effect, proceed to the Main cyCombine workflow (Python port) tutorial to run the correction pipeline.