Pairwise Comparisons with stat_pwc¶
This vignette demonstrates how to use stat_pwc to add pairwise comparison p-values to plots, similar to ggpubr’s ``geom_pwc` <https://rpkgs.datanovia.com/ggpubr/reference/geom_pwc.html>`__.
stat_pwc performs pairwise statistical tests (t-test, Wilcoxon) between groups and displays the results as bracket annotations with p-values. It supports:
All pairwise combinations
Comparisons against a reference group
Explicit comparison pairs
Multiple p-value adjustment methods (Bonferroni, Holm, BH, etc.)
Various label formats (p-value, significance stars)
Setup¶
[1]:
import numpy as np
import pandas as pd
from plotnine import (
aes,
geom_boxplot,
geom_jitter,
geom_point,
ggplot,
labs,
scale_y_continuous,
theme_minimal,
facet_wrap,
)
from plotnine_extra import stat_pwc
Data Preparation¶
We use a dataset inspired by R’s ToothGrowth dataset, which records tooth length (len) across different supplement types (supp) and dose levels (dose).
[2]:
np.random.seed(42)
# Create a ToothGrowth-like dataset
df = pd.DataFrame({
"dose": np.tile(np.repeat(["D0.5", "D1", "D2"], 10), 2),
"supp": np.repeat(["OJ", "VC"], 30),
"len": np.concatenate([
# OJ groups
np.random.normal(13, 4, 10), # OJ, dose 0.5
np.random.normal(23, 3, 10), # OJ, dose 1
np.random.normal(26, 2, 10), # OJ, dose 2
# VC groups
np.random.normal(8, 3, 10), # VC, dose 0.5
np.random.normal(17, 4, 10), # VC, dose 1
np.random.normal(26, 3, 10), # VC, dose 2
]),
})
df.head()
[2]:
| dose | supp | len | |
|---|---|---|---|
| 0 | D0.5 | OJ | 14.986857 |
| 1 | D0.5 | OJ | 12.446943 |
| 2 | D0.5 | OJ | 15.590754 |
| 3 | D0.5 | OJ | 19.092119 |
| 4 | D0.5 | OJ | 12.063387 |
Basic Usage: All Pairwise Comparisons¶
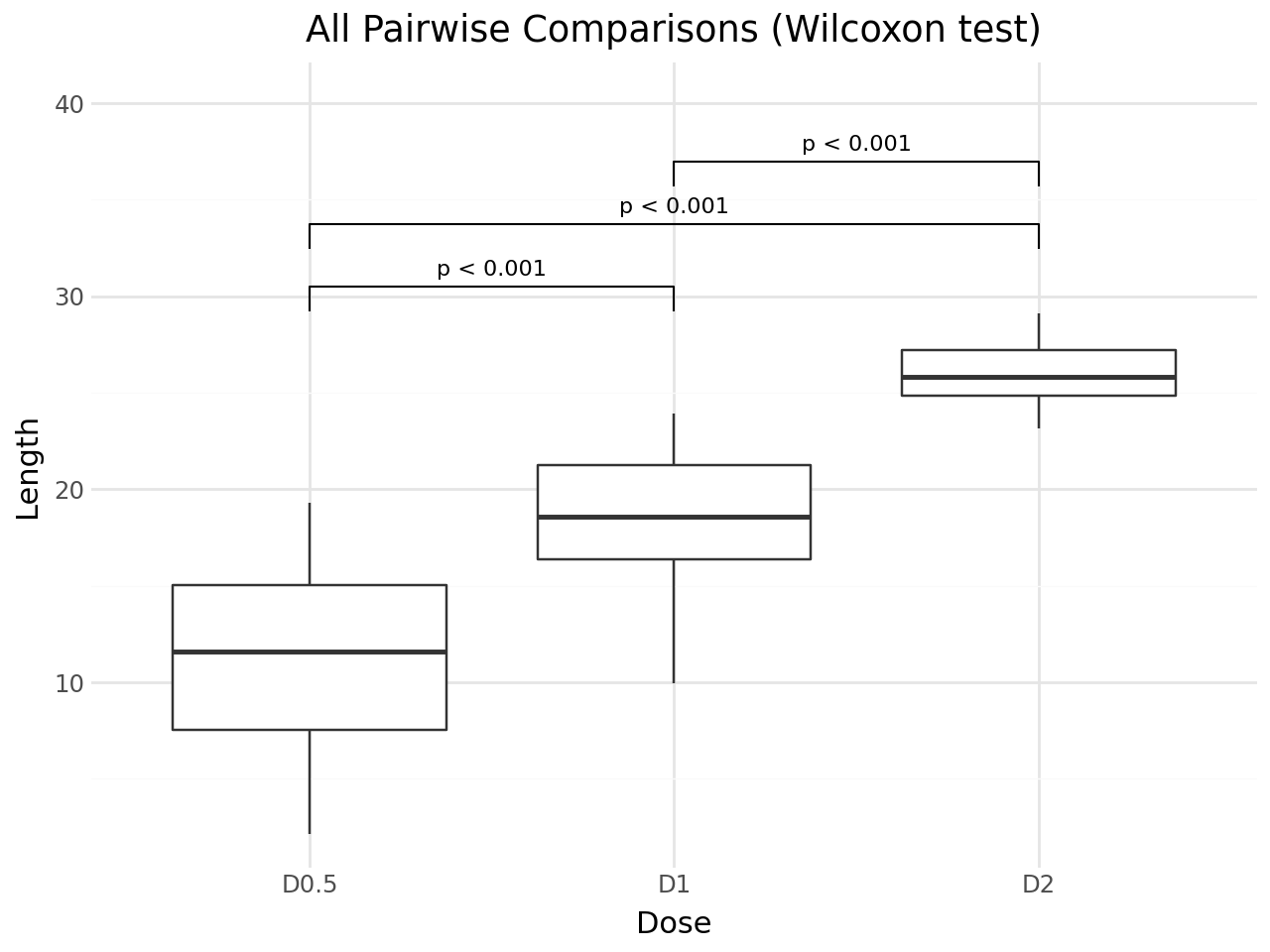
By default, stat_pwc performs all pairwise comparisons between x-axis groups using the Wilcoxon rank-sum test (Mann-Whitney U). The default geom is geom_bracket, which draws brackets with p-value labels.
[3]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot()
+ stat_pwc()
+ scale_y_continuous(expand=(0.05, 0, 0.15, 0))
+ labs(
title="All Pairwise Comparisons (Wilcoxon test)",
x="Dose",
y="Length",
)
+ theme_minimal()
)
[3]:
Using t-test Instead of Wilcoxon¶
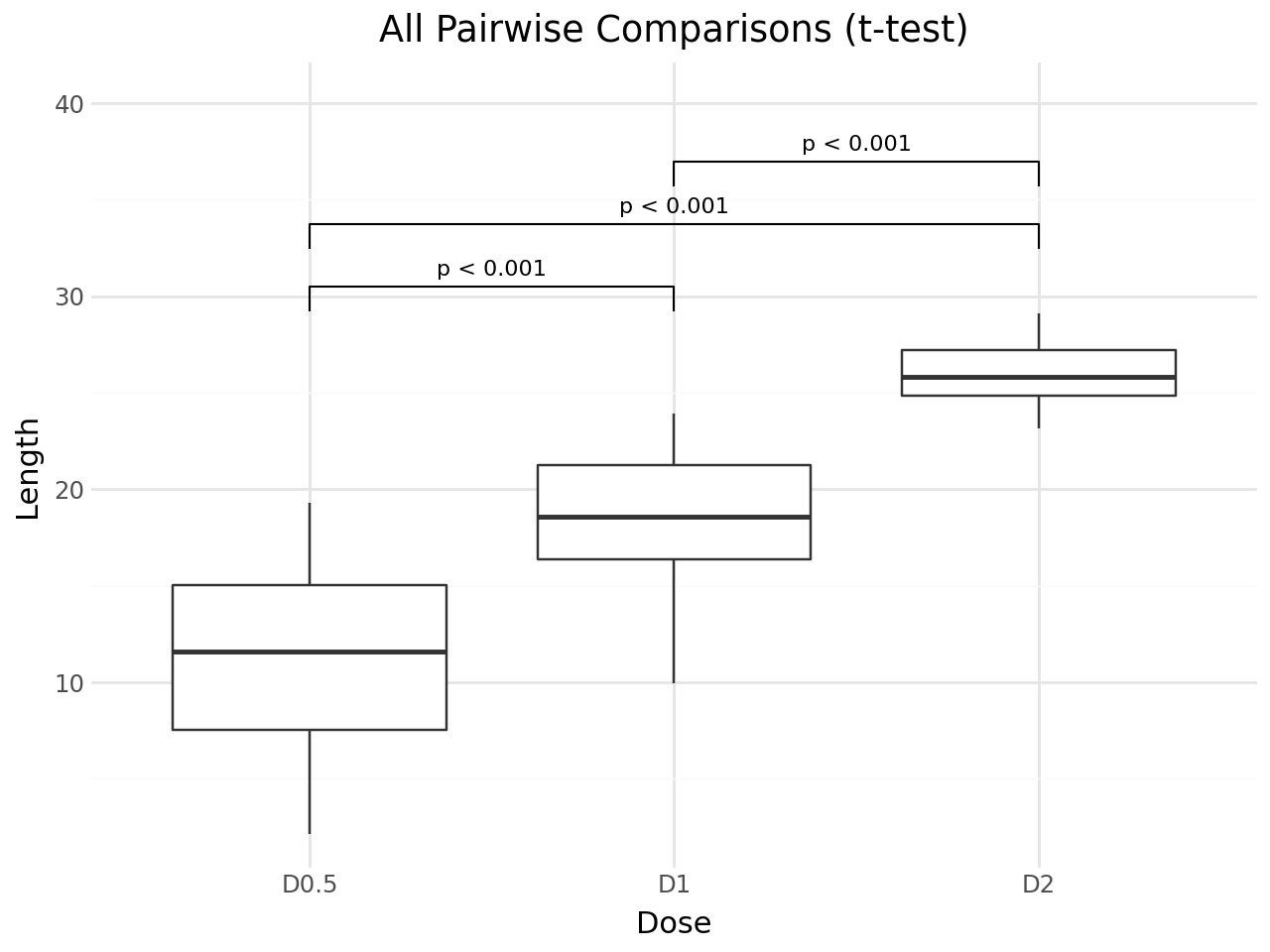
Set method="t.test" to use the independent samples t-test.
[4]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot()
+ stat_pwc(method="t.test")
+ scale_y_continuous(expand=(0.05, 0, 0.15, 0))
+ labs(
title="All Pairwise Comparisons (t-test)",
x="Dose",
y="Length",
)
+ theme_minimal()
)
[4]:
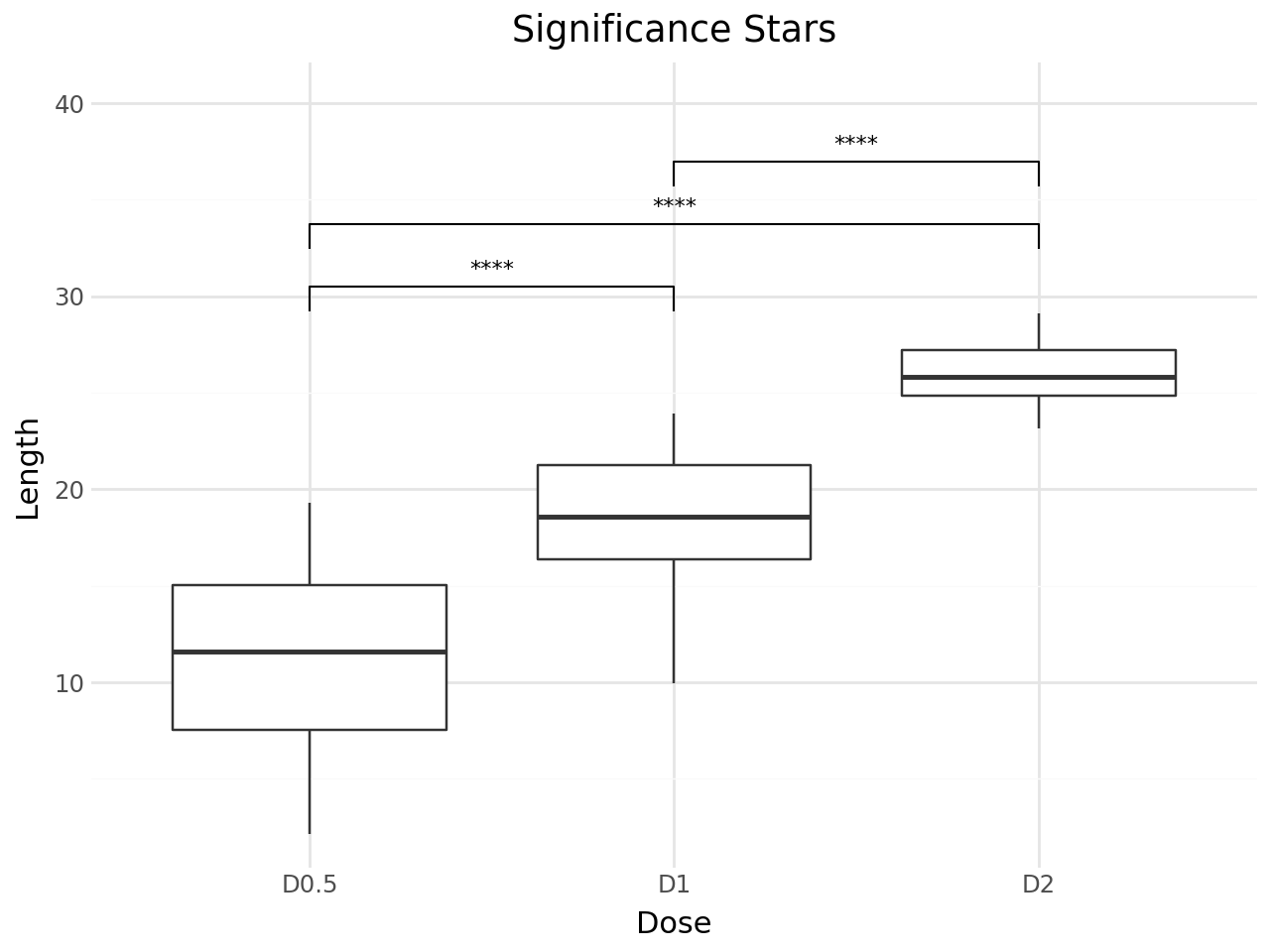
Significance Stars¶
Use label="p.signif" to display significance stars instead of p-values. The convention is:
Symbol |
Meaning |
|---|---|
|
p > 0.05 |
|
p ≤ 0.05 |
|
p ≤ 0.01 |
|
p ≤ 0.001 |
|
p ≤ 0.0001 |
[5]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot()
+ stat_pwc(label="p.signif")
+ scale_y_continuous(expand=(0.05, 0, 0.15, 0))
+ labs(
title="Significance Stars",
x="Dose",
y="Length",
)
+ theme_minimal()
)
[5]:
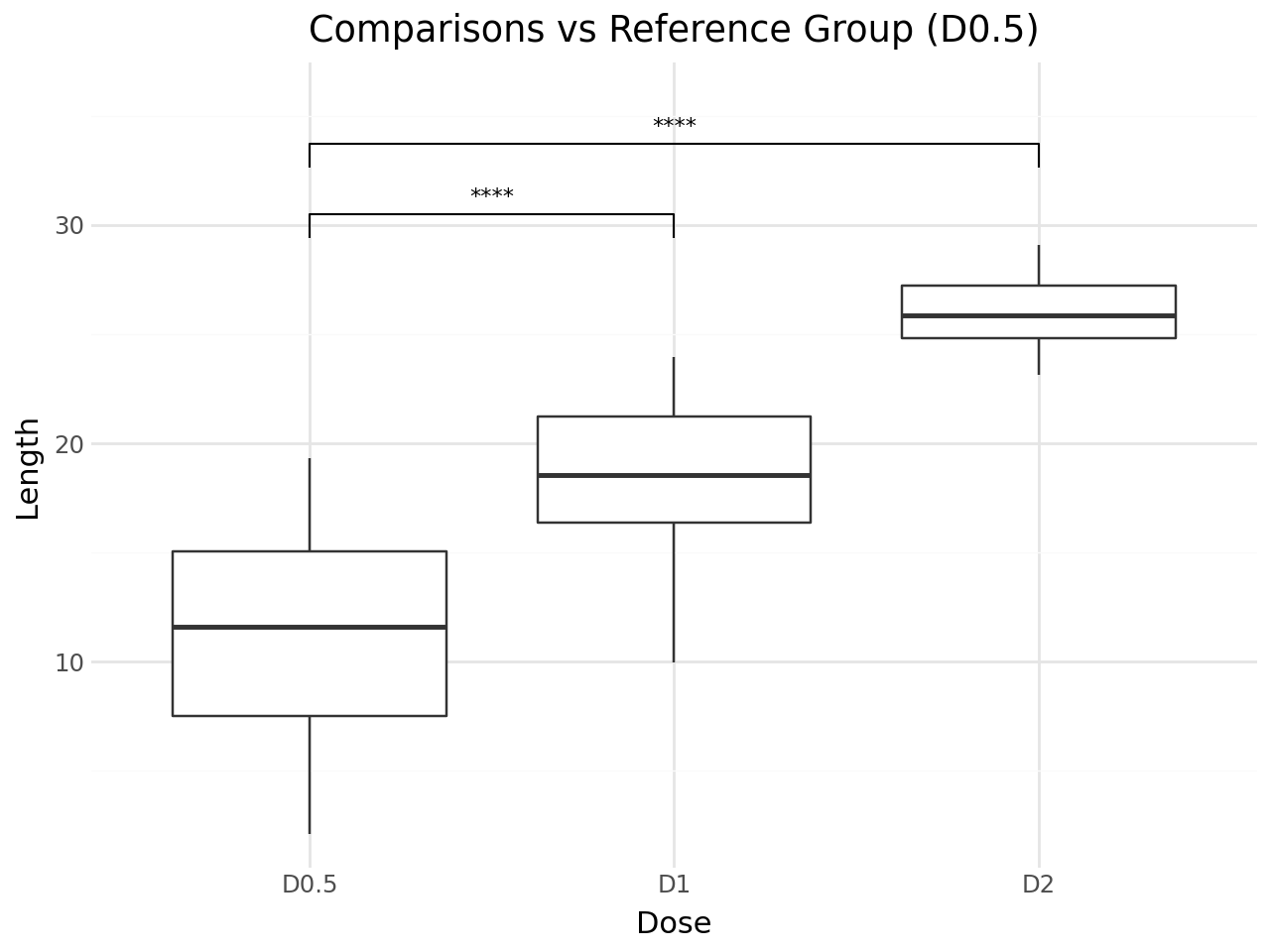
Comparisons Against a Reference Group¶
Use ref_group to compare each group against a reference. This is useful for comparing treatment groups against a control.
[6]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot()
+ stat_pwc(
ref_group="D0.5",
label="p.signif",
method="t.test",
)
+ scale_y_continuous(expand=(0.05, 0, 0.12, 0))
+ labs(
title='Comparisons vs Reference Group (D0.5)',
x="Dose",
y="Length",
)
+ theme_minimal()
)
[6]:
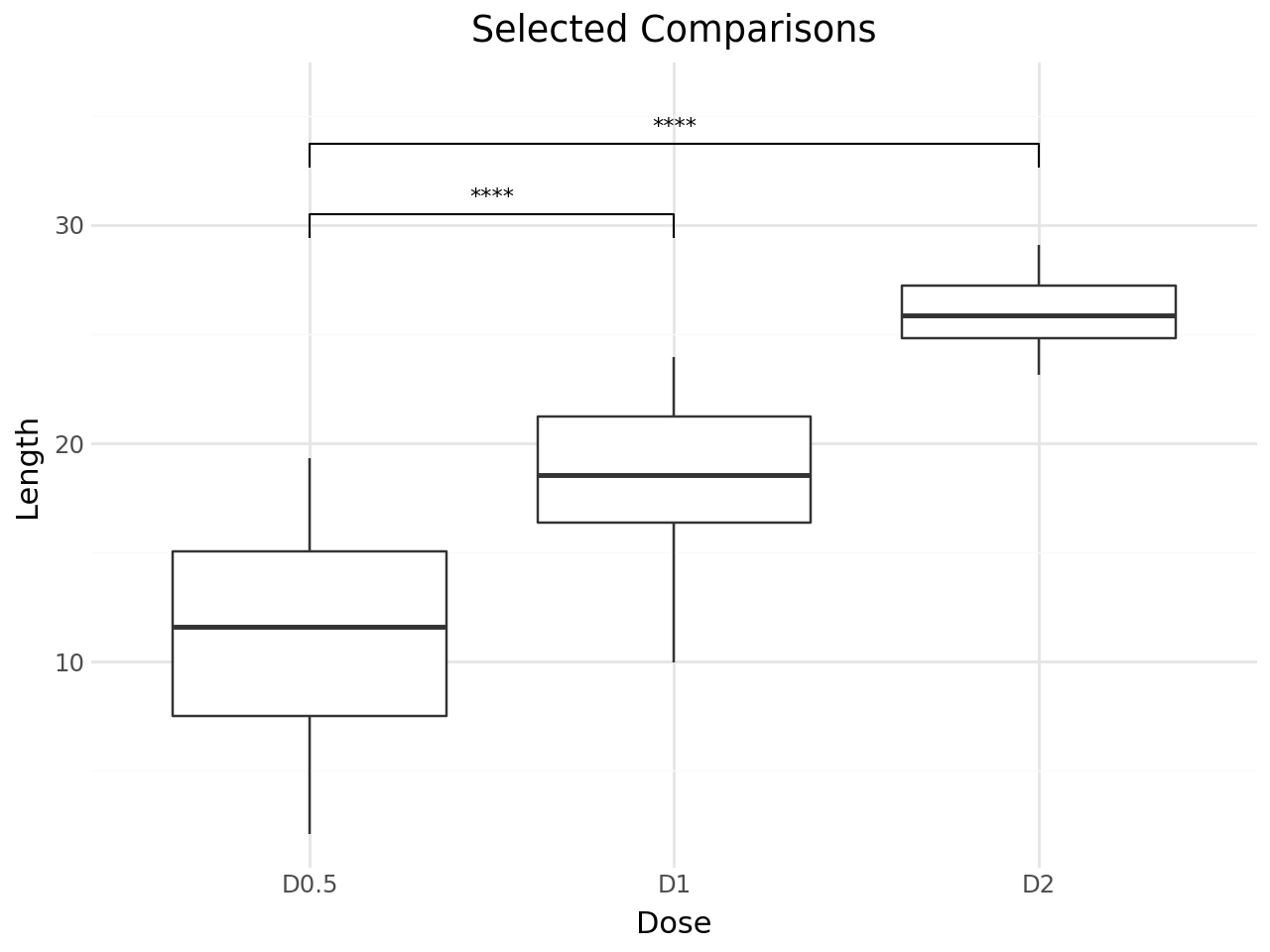
Explicit Comparison Pairs¶
Specify exactly which pairs to compare with the comparisons parameter.
[7]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot()
+ stat_pwc(
comparisons=[("D0.5", "D1"), ("D0.5", "D2")],
label="p.signif",
)
+ scale_y_continuous(expand=(0.05, 0, 0.12, 0))
+ labs(
title="Selected Comparisons",
x="Dose",
y="Length",
)
+ theme_minimal()
)
[7]:
P-value Adjustment¶
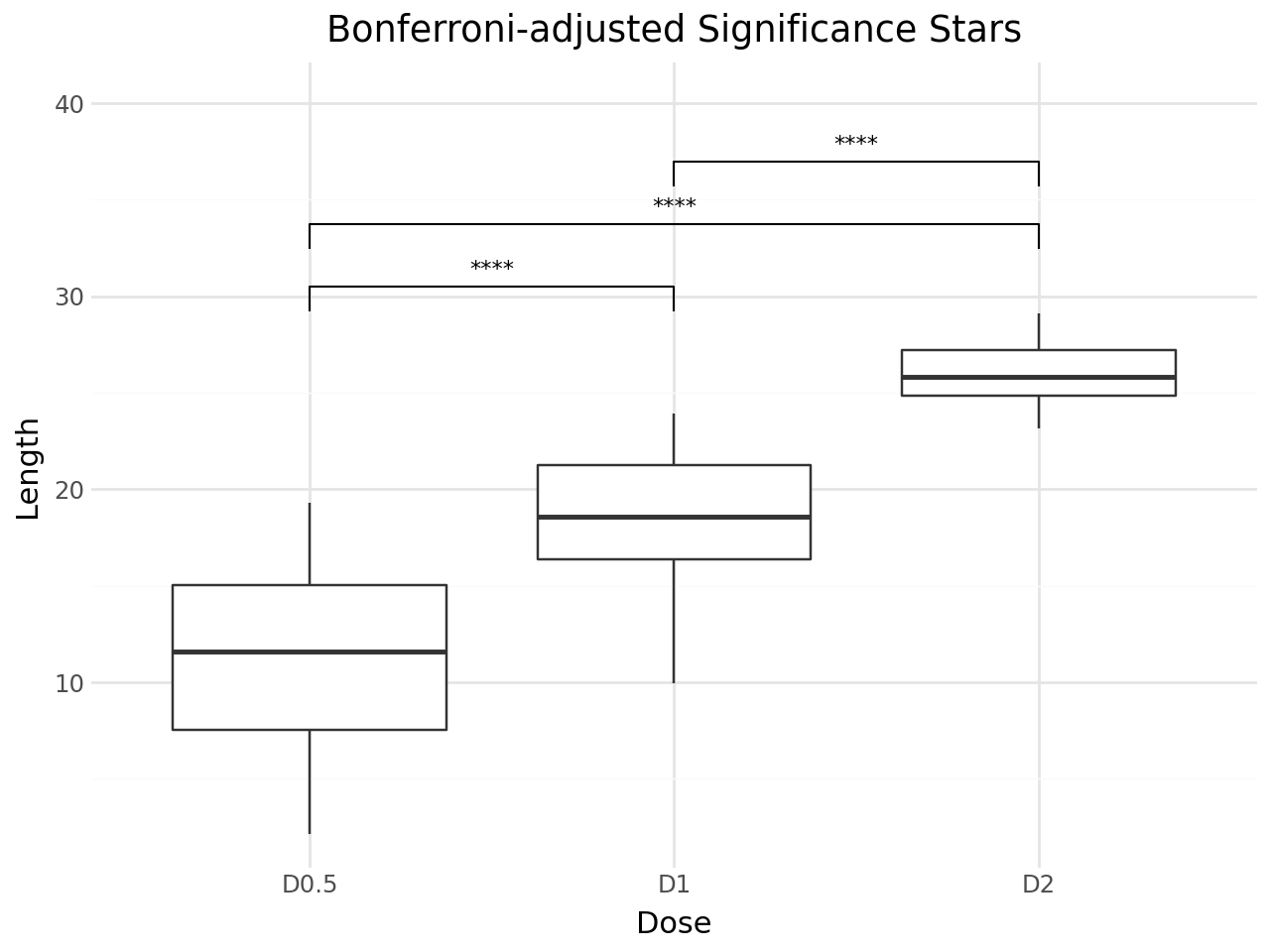
stat_pwc adjusts p-values for multiple comparisons by default using the Holm method. You can change the adjustment method with p_adjust_method.
Use label="p.adj.format" to display adjusted p-values, or label="p.adj.signif" for adjusted significance stars.
[8]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot()
+ stat_pwc(
label="p.adj.signif",
p_adjust_method="bonferroni",
method="t.test",
)
+ scale_y_continuous(expand=(0.05, 0, 0.15, 0))
+ labs(
title="Bonferroni-adjusted Significance Stars",
x="Dose",
y="Length",
)
+ theme_minimal()
)
[8]:
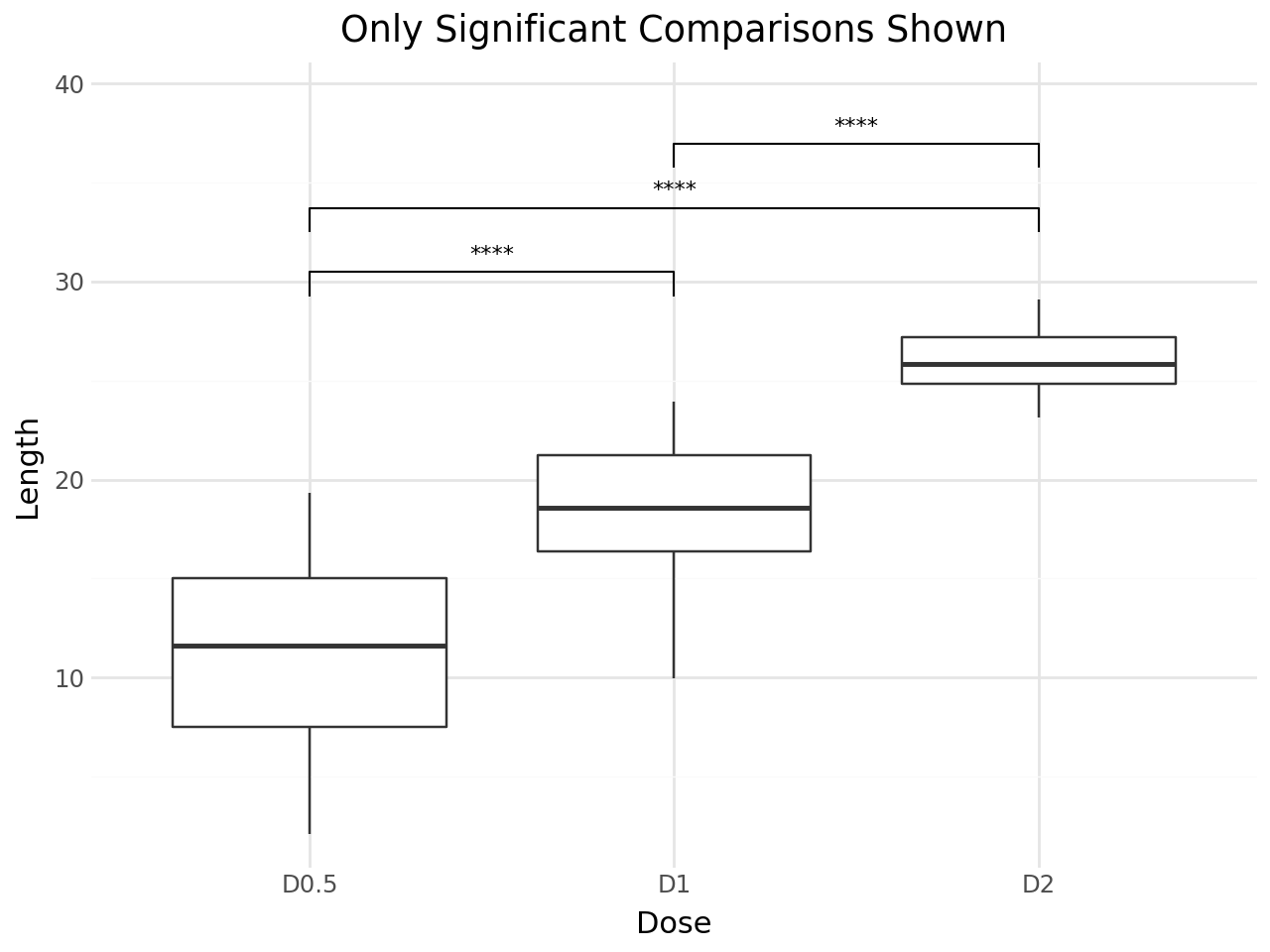
Hiding Non-significant Comparisons¶
Set hide_ns=True to remove non-significant comparisons (p > 0.05) from the plot.
[9]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot()
+ stat_pwc(
label="p.signif",
hide_ns=True,
)
+ scale_y_continuous(expand=(0.05, 0, 0.12, 0))
+ labs(
title="Only Significant Comparisons Shown",
x="Dose",
y="Length",
)
+ theme_minimal()
)
[9]:
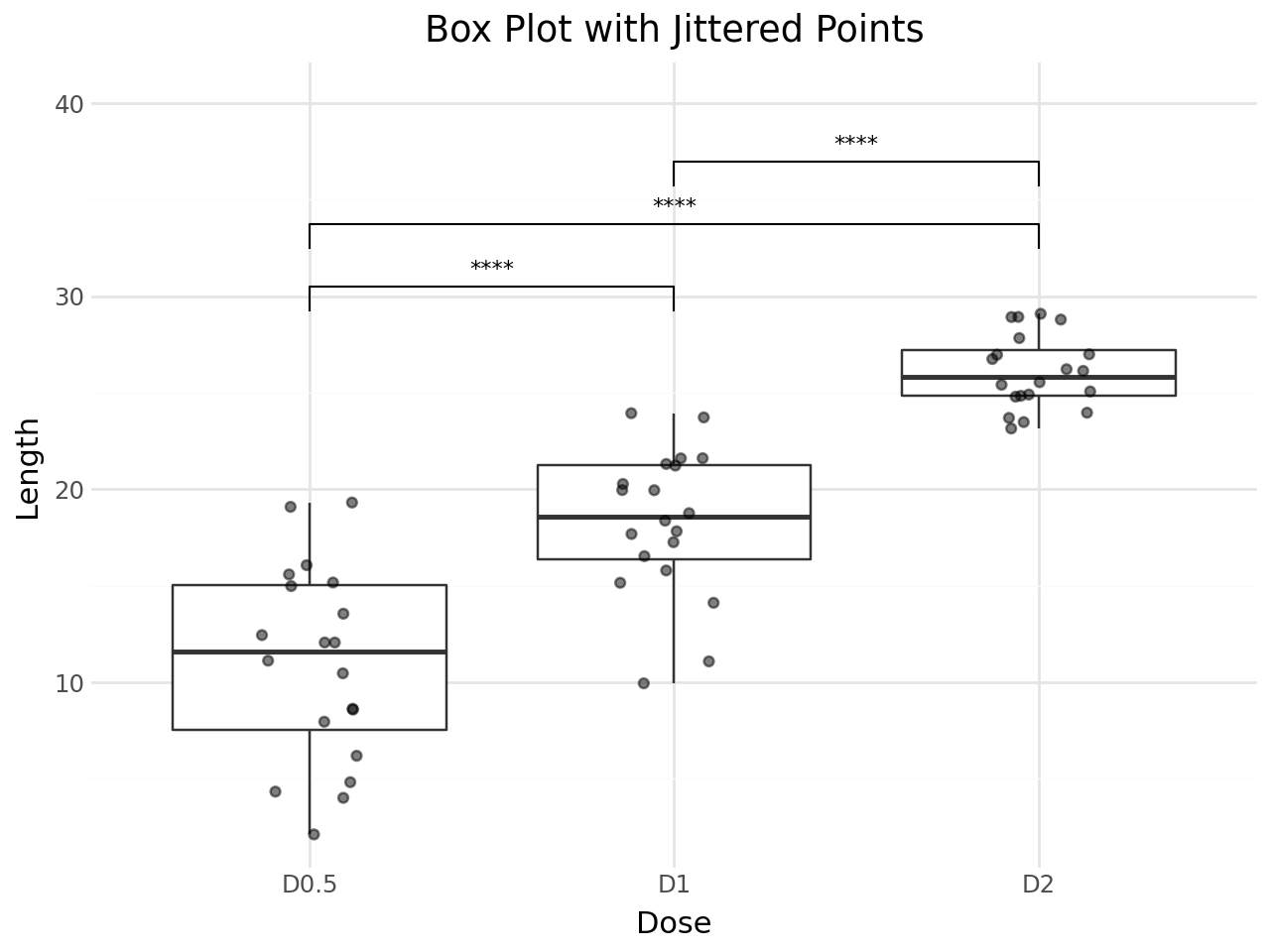
Box Plot with Jittered Points¶
stat_pwc works with any geom. Here we combine box plots with jittered data points for a richer visualization.
[10]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot(outlier_shape="")
+ geom_jitter(width=0.15, alpha=0.5)
+ stat_pwc(
method="t.test",
label="p.signif",
)
+ scale_y_continuous(expand=(0.05, 0, 0.15, 0))
+ labs(
title="Box Plot with Jittered Points",
x="Dose",
y="Length",
)
+ theme_minimal()
)
[10]:
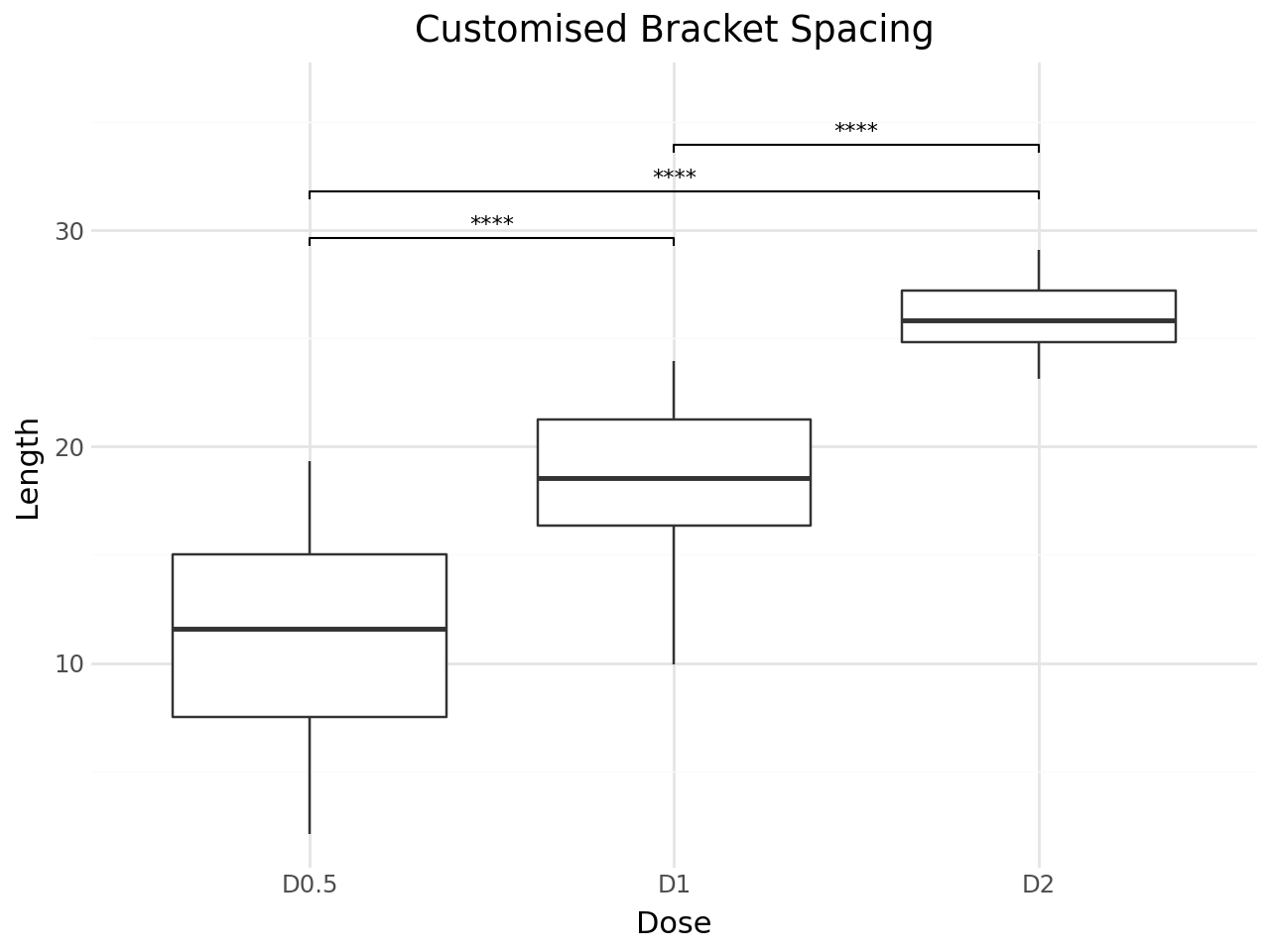
Customising Bracket Appearance¶
You can adjust the bracket layout with:
step_increase– gap between stacked brackets (fraction of y-range)bracket_nudge_y– vertical offset for all bracketstip_length– length of bracket tips
[11]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot()
+ stat_pwc(
label="p.signif",
step_increase=0.08,
bracket_nudge_y=0.02,
tip_length=0.01,
)
+ scale_y_continuous(expand=(0.05, 0, 0.12, 0))
+ labs(
title="Customised Bracket Spacing",
x="Dose",
y="Length",
)
+ theme_minimal()
)
[11]:
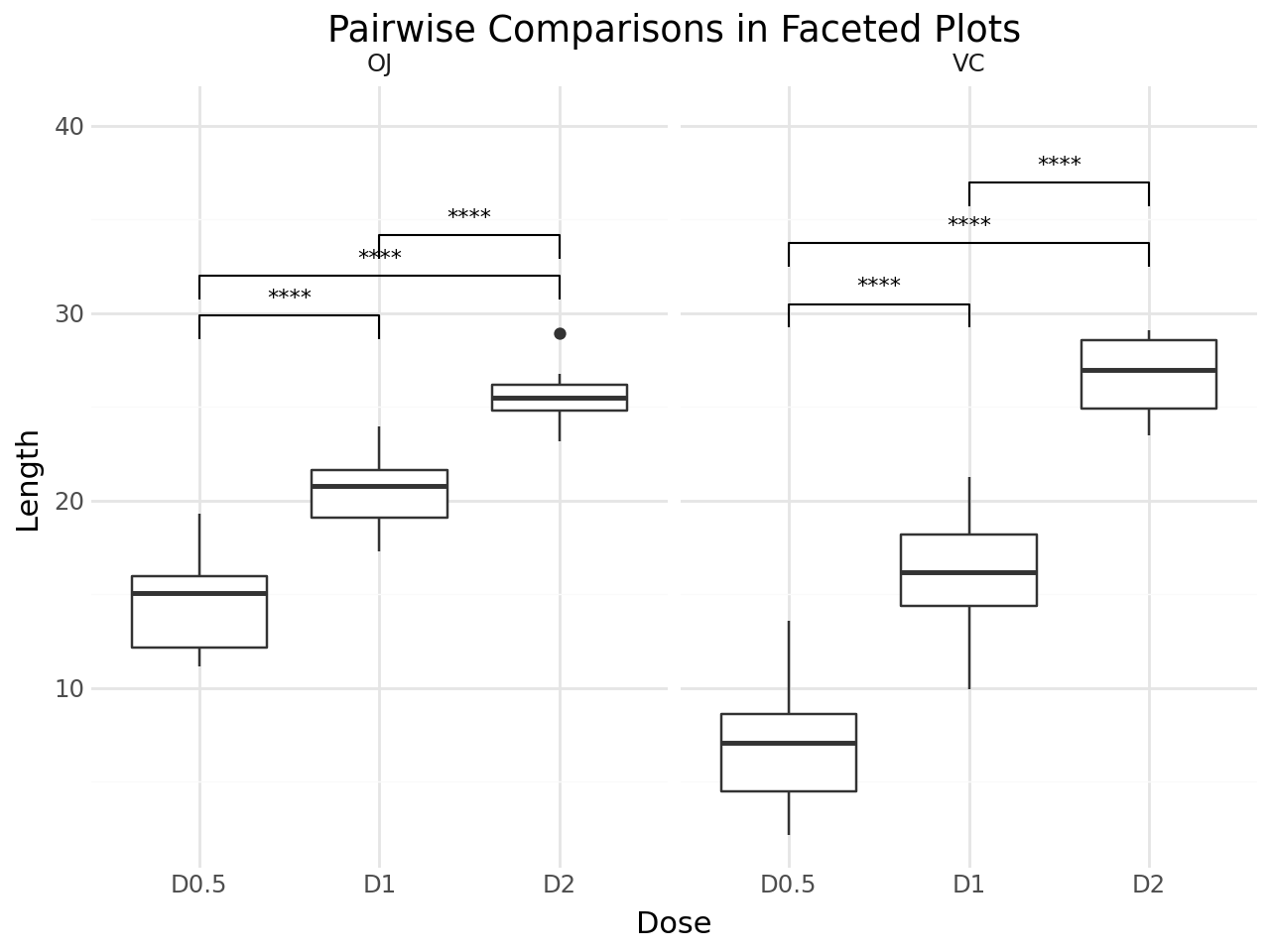
Faceted Plots¶
stat_pwc works with faceted plots. Each facet panel gets its own set of pairwise comparisons.
[12]:
(
ggplot(df, aes(x="dose", y="len"))
+ geom_boxplot()
+ stat_pwc(
label="p.signif",
method="t.test",
)
+ facet_wrap("supp")
+ scale_y_continuous(expand=(0.05, 0, 0.15, 0))
+ labs(
title="Pairwise Comparisons in Faceted Plots",
x="Dose",
y="Length",
)
+ theme_minimal()
)
[12]:
Available Label Formats¶
|
Description |
|---|---|
|
Formatted raw p-value (default) |
|
Significance symbols (ns, *, **, ***, ****) |
|
Formatted adjusted p-value |
|
Adjusted significance symbols |
|
Raw p-value with significance symbol |
Available P-value Adjustment Methods¶
|
Description |
|---|---|
|
Holm (default, step-down) |
|
Bonferroni |
|
Hochberg |
|
Benjamini-Hochberg (FDR) |
|
Benjamini-Yekutieli |
|
Hommel |
|
No adjustment |
Summary¶
stat_pwc makes it easy to add statistical comparison annotations to any plotnine chart. Key features:
Automatic pairwise testing between all groups, or against a reference group
Multiple test methods: Wilcoxon rank-sum (default) and t-test
P-value adjustment for multiple comparisons
Flexible labelling: p-values, significance stars, or both
Bracket customisation: spacing, tips, and nudging
Works with facets — each panel gets its own comparisons
For more details, see the API reference.